-
If you have not already done so, request a user account/allocation on the Center for Research Computing (CRC) cluster by filling out the required information on this page
-
If your computer is not connected to the Pitt network (e.g. you are working from home or on a trip), or you are working from a laptop that is connected to the UPMC network, make sure you setup Pitt SSLVPN, so that you can communicate with the Center for Research Computing (CRC) cluster (galaxy servers are using HTC cluster). Make sure that "Server URL" (4) is sremote.pitt.edu, and "Please select a Role" (14) is Firewall-SAM-USERS-Pulse. Note that there are many different VPN roles. Only Firewall-SAM-USERS-Pulse role can connect to CRC clusters. If your VPN is installed by system administrators and you are not sure what role is used, open Pulse Secure, and click + sign and follow the instructions.
-
The Server host is http://galaxy.crc.pitt.edu:8080. Fill in your Pitt username and password. Please note that username is case sensitive and all letters are in lowercase. Refer to the image below for an example of how the settings in this box should look:
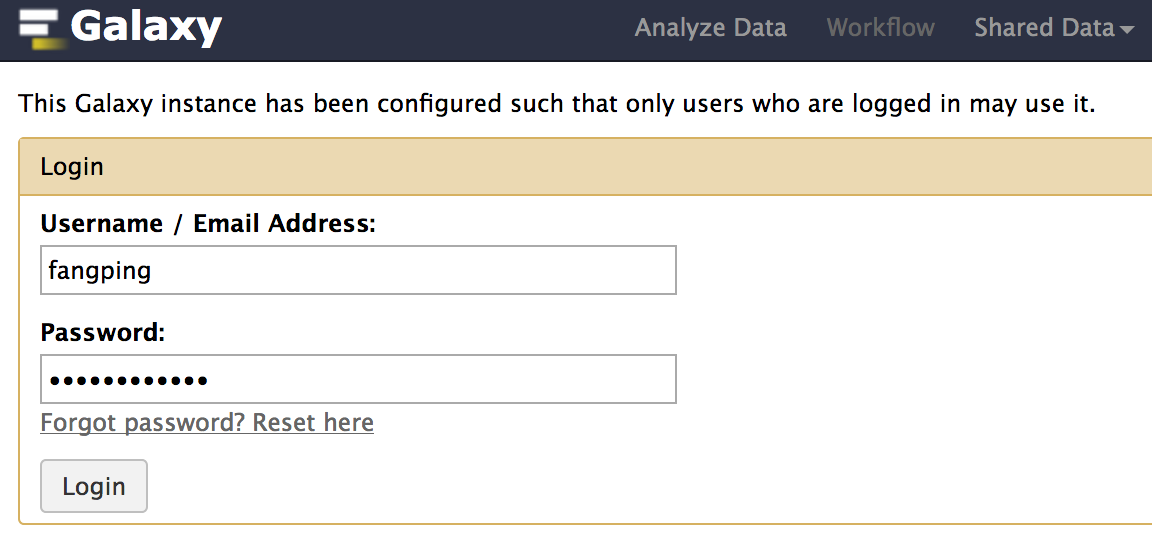
-
You can load data from your local computer to galaxy history using the "Get Data > Update File" tool. To load large files, you can use galaxy data libraries.
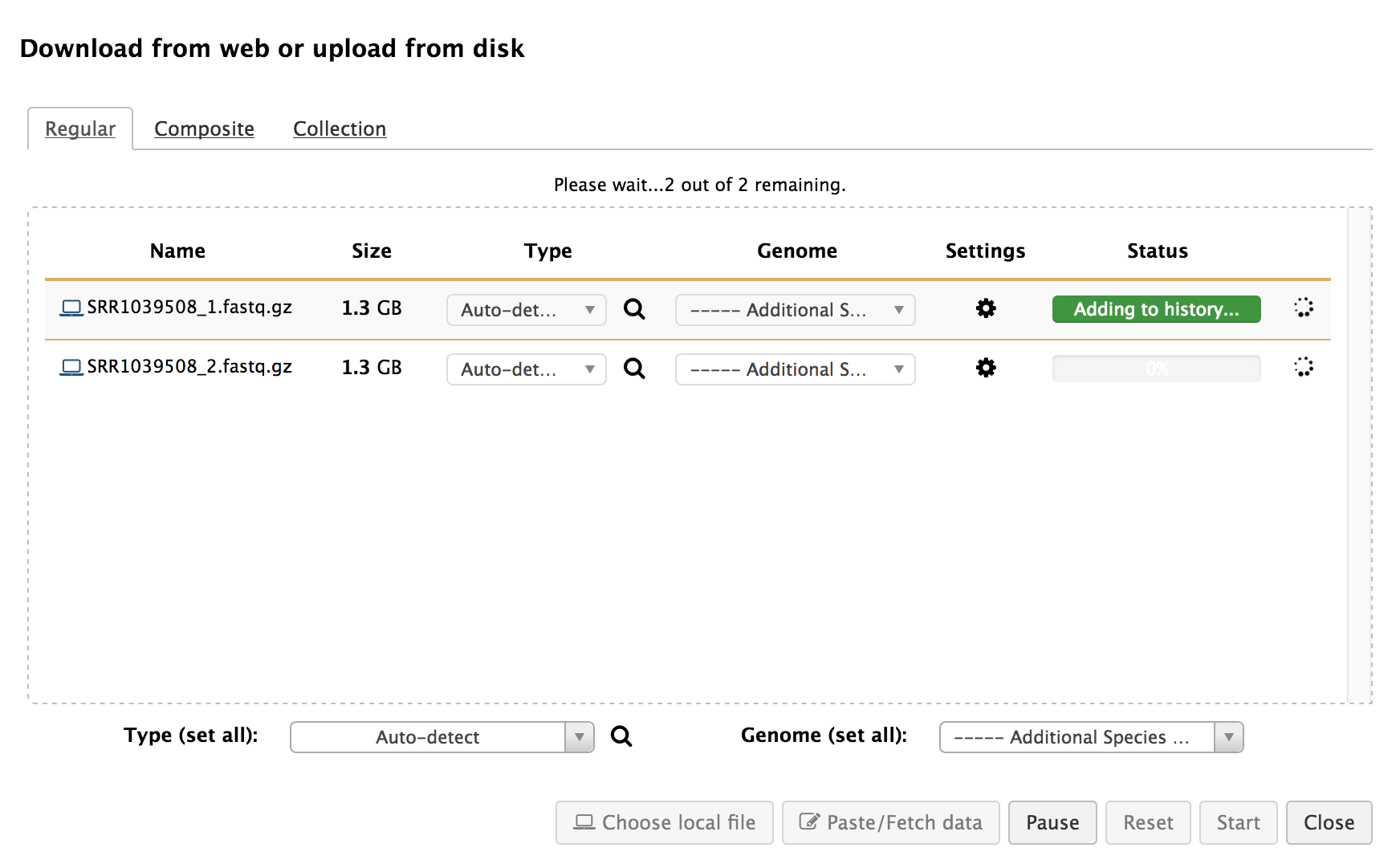
-
Running an analysis operates in choosing the tool from left panel and filling in the parameters. The following image shows using the [NGS: RNA Analysis >] HISAT2 tool to map RNA-seq reads to the human hg38 genmome. Every time a tool is run, one or more datasets are created in the user's history.
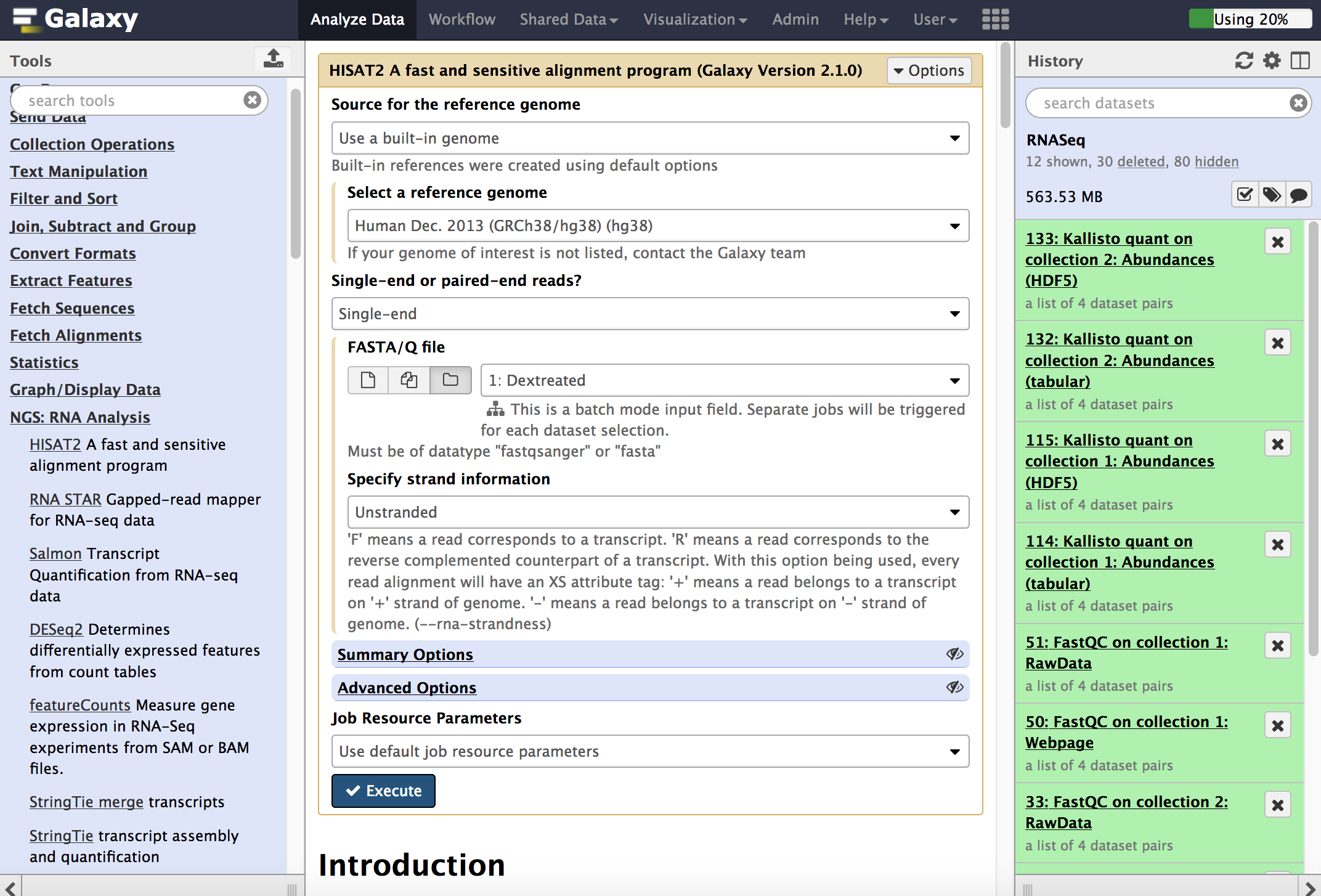
-
Once you start a job, it will be running on HTC cluster, except the tools under "Get Data", which will be run on galaxy server computer. You will see the status of your jobs on the history.
-
You can submit a CRC help ticket to request a galaxy data libraries. You can use standard data transferring tools, such as rsync, FileZilla, Cyberduck to transfer data to your galaxy data libraries.
-
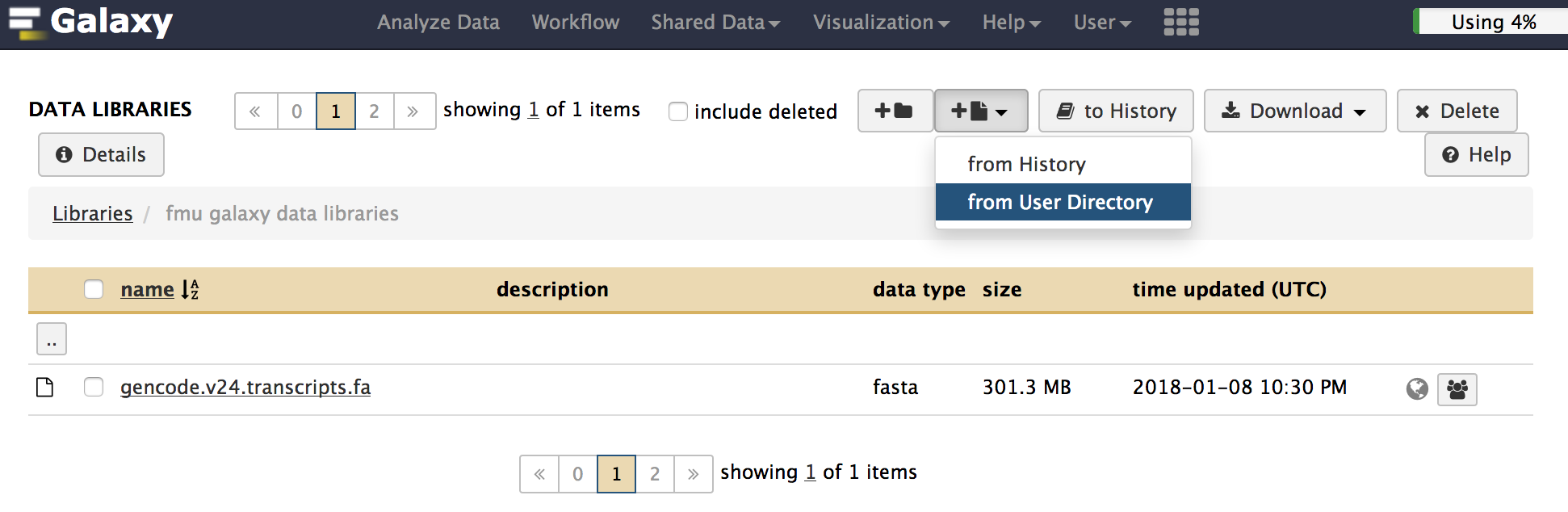
To transfer data from your galaxy data libraries to History, click Shared Data -> Data Libraries to go to your data libraries. Then Add Datasets to Current Folder -> from User Directory. Note that data transferring needs time, and you may see that your files are 0 bytes for a while.
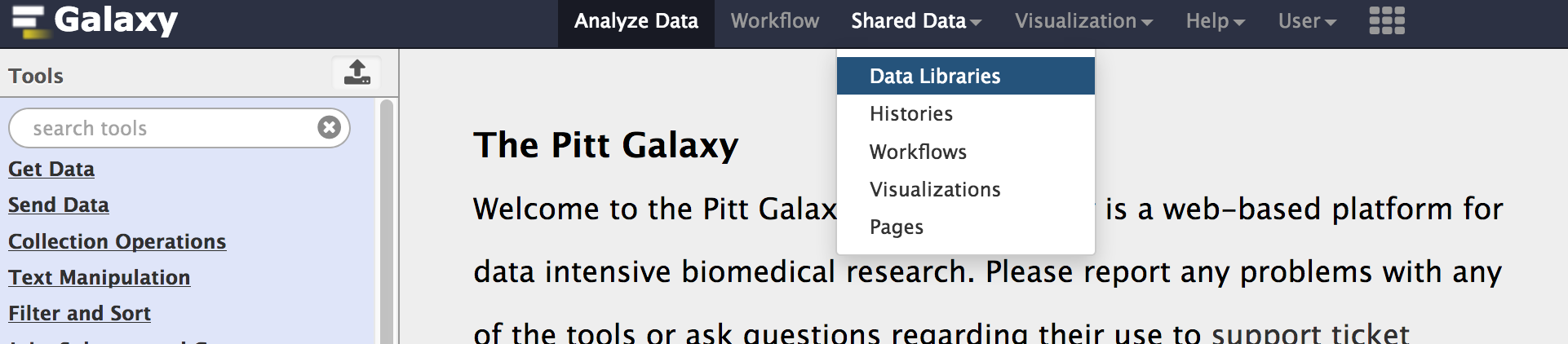
-
Vast number of good and well-implemented tools for Galaxy is available through the Galaxy ToolShed. You can submit a CRC help ticket to request installation of them into this Galaxy instance.
-
Users are assigned default disk quota of 1 TB.
-
Galaxy data are assigned on /zfs1. Note that this /zfs1 is not backed up, so you will need to be diligent and back up to your own personal data.
-
If you have any problems with this procedure, please submit a CRC support ticket online (login required)
